PDBsum Proprietary.
3D Protein Structure and Ligand complex repository, analysis and communication system.
PDBsum Proprietary
Empower collaboration between structural biologists as well as computational & medicinal chemists.
Install and run behind your own firewall
Facilitate analysis of proprietary 3D structures and ligand complexes.
PDBsumProprietary teases out key points of a protein complex for intial analysis and avoids the requirement for specialist 3D viewing software and knowledge.
Repository
Add all 3D structures determined in-house, by either x-ray or NMR experiments or predictive homology models. Preview the full functionality of PDBsumProprietary by simply looking at the public version of PDBsum which processes all public PDB structures at the European Bioinformatics Institute (see links in examples below).
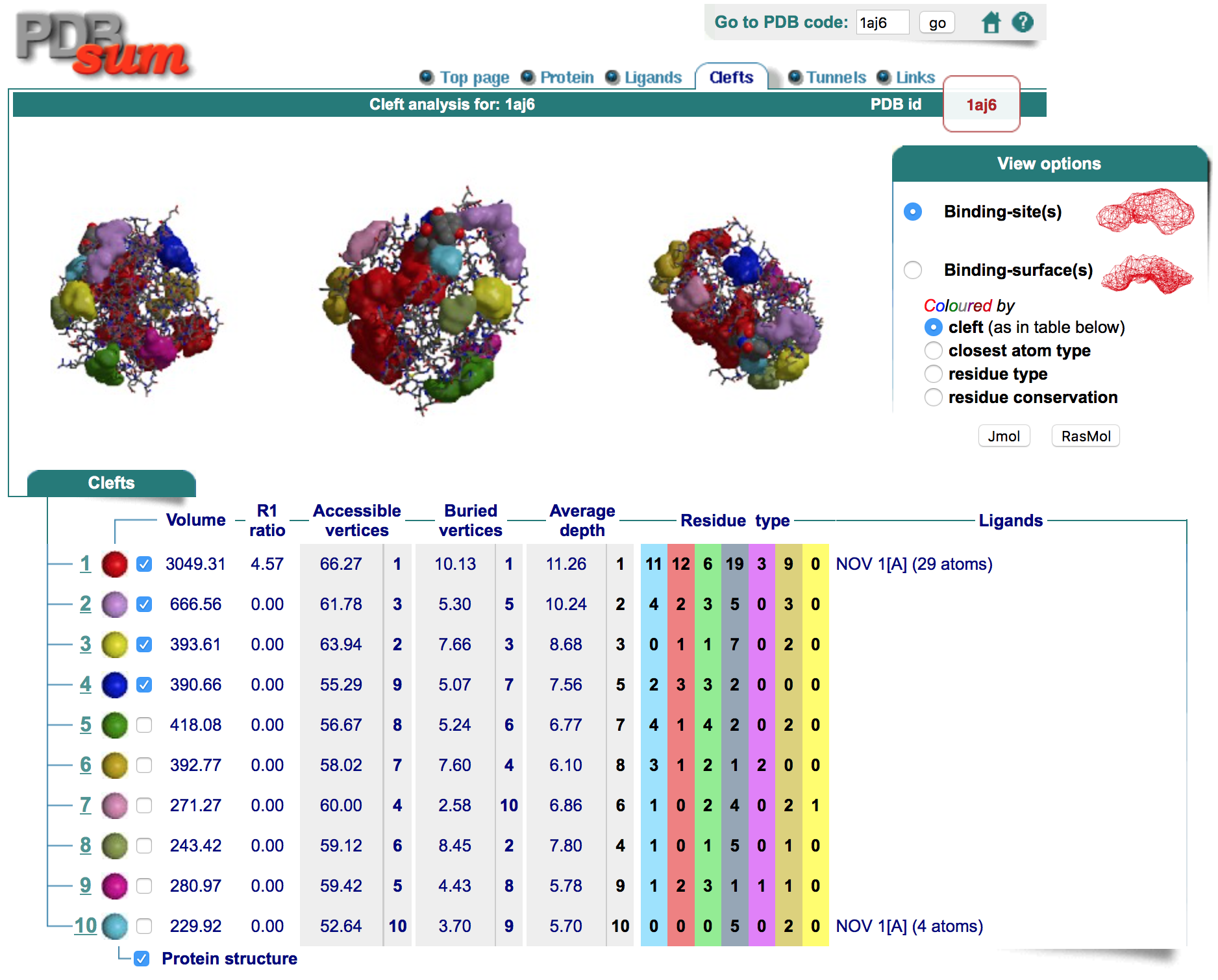
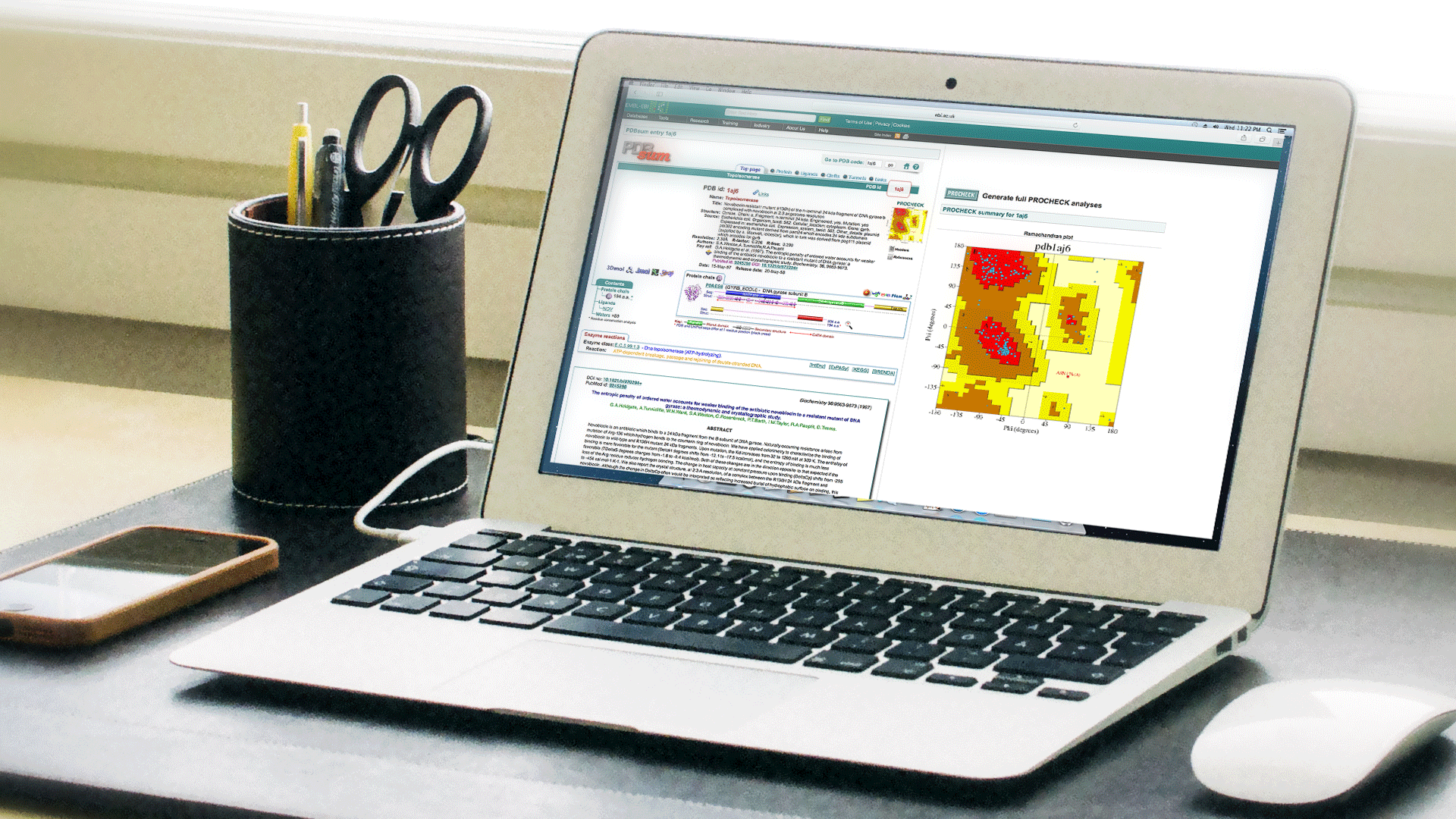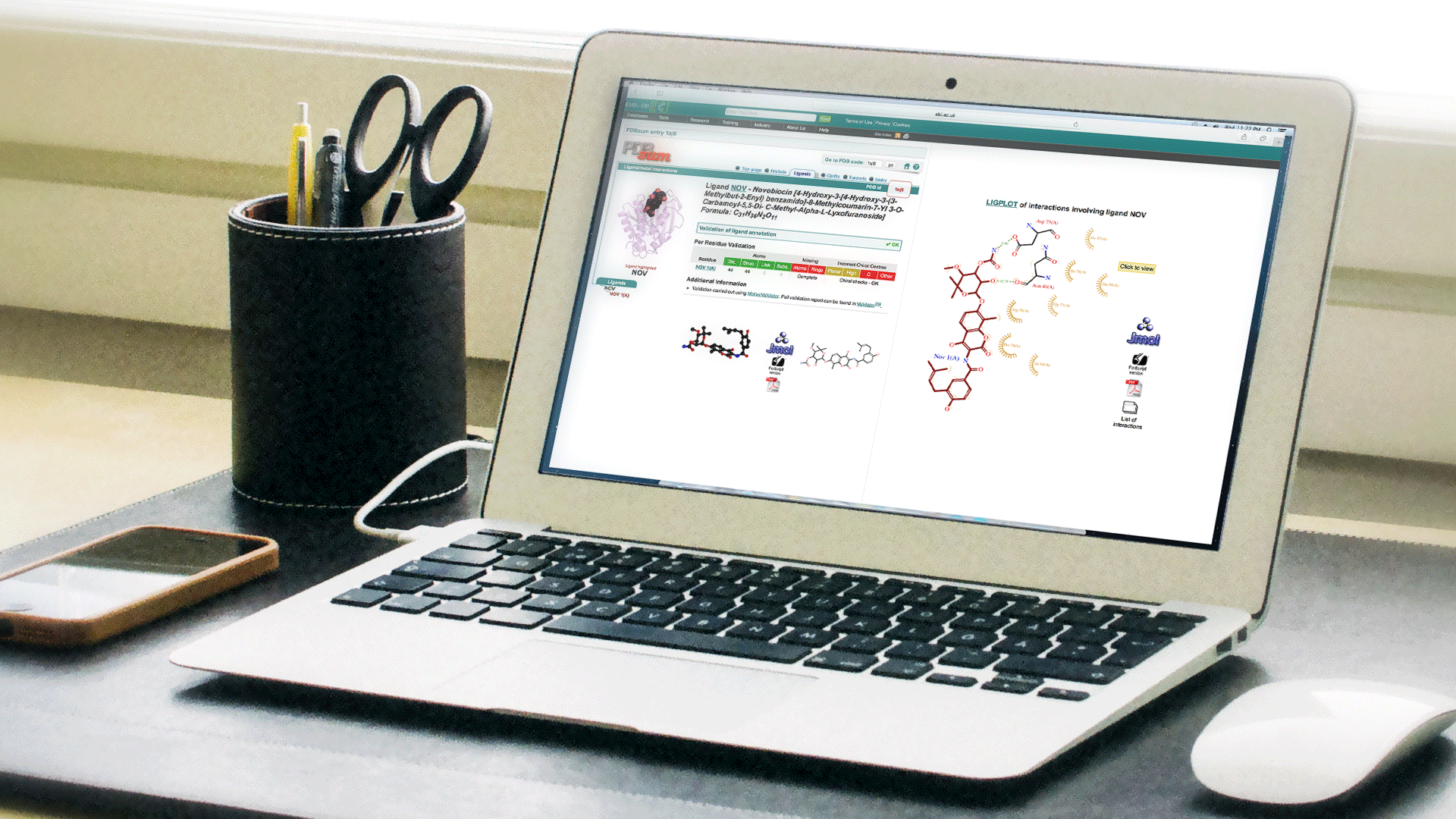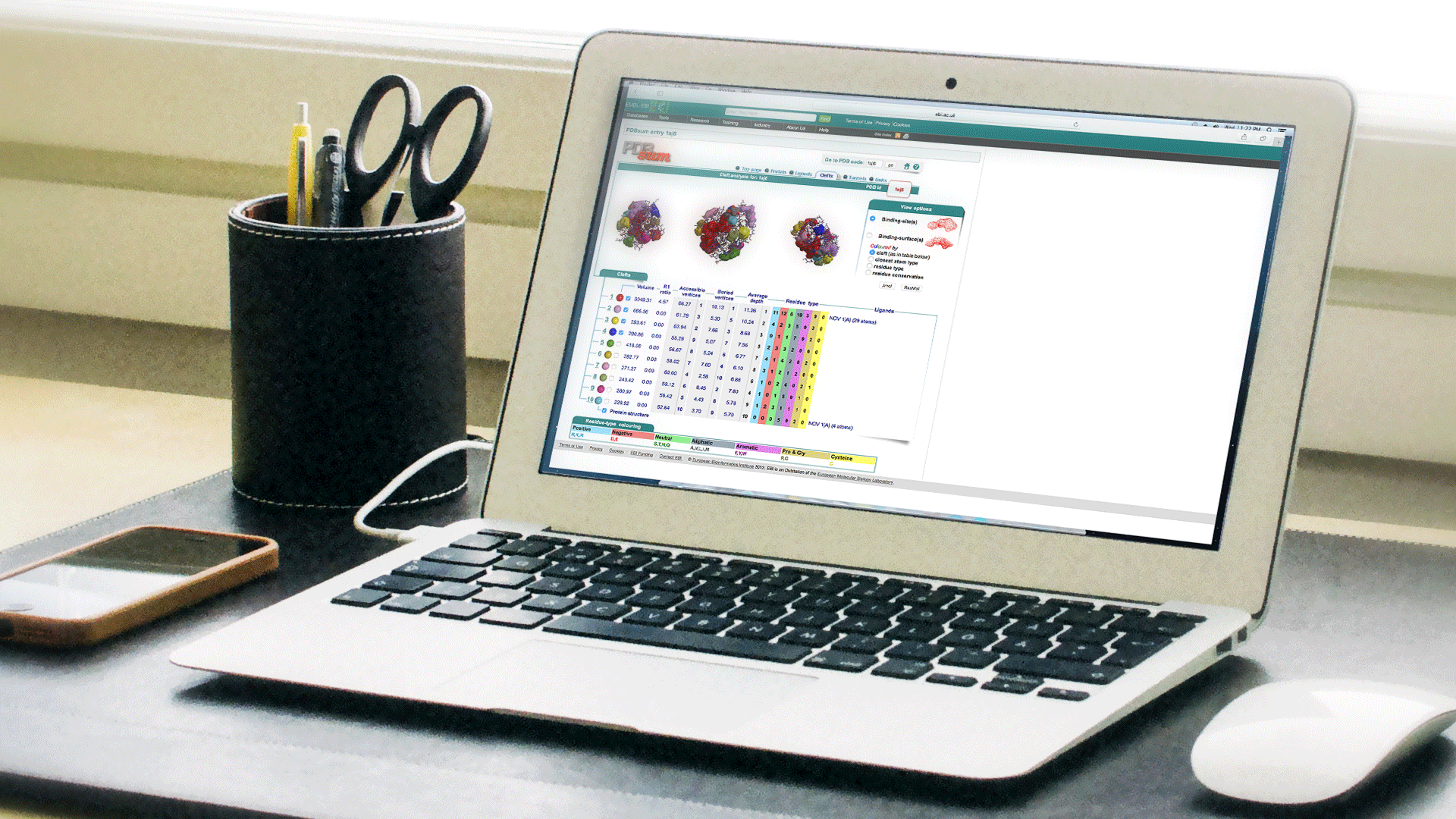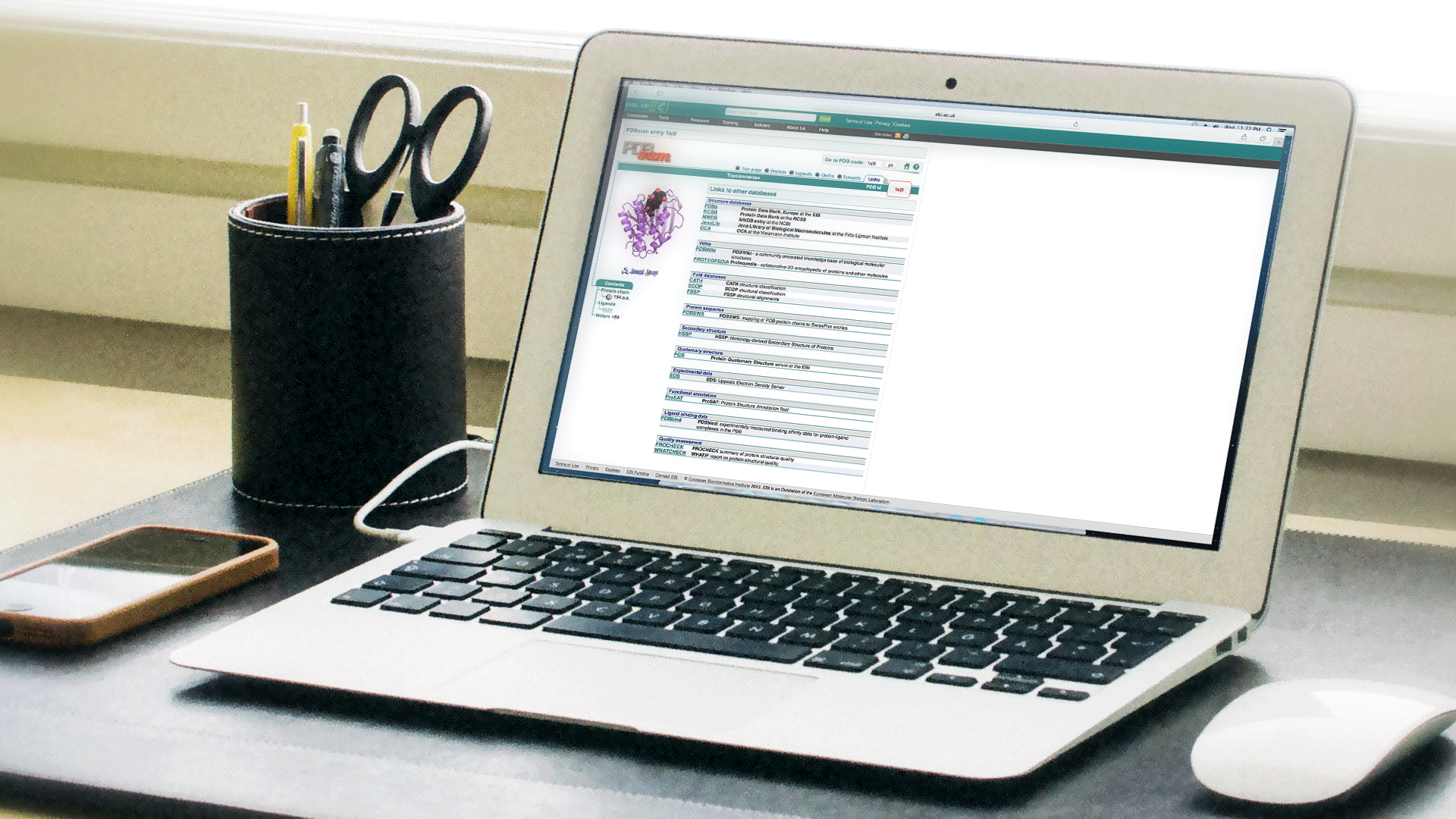
Top Page
Simple Overview of Structure including detailed quality interpretation through web implemented Procheck and Ramachandran plot.
Data Input
Data can be input in two ways.
Completed structures are introduced through a pipeline script that is excecuted as a cron job at regular intervals that suits your work requirements. Alternatively, if you want to take a quick look at an interim structure file, perhaps during the refinement process or to check whether a homology model makes sense, you can use the Generate function will allow just you to see a temporary set of web pages. These temporary pages are then cleared according to a schedule chosen by your group.
3D Viewing
Although the focus of this web site is simple 2D viewing, various features can also the 3D enabled using JMol or RasMol for quick analysis.
Public Version of PDBsum
A quick reminder that the public version of PDBsum is hosted at the European Bioinformatics Institute in Prof. Janet Thornton’s group.